4.1 Environment and Configuration
The present study inherits the technical framework defined by the reference work in Marín
et al. (
2020). Consequently, the GrC model is simulated with the neural simulation tool NEST (Plesser
et al.,
2015), in its version 2.14.0, and using the 4th-order Runge-Kutta-Fehlberg solver with adaptive step-size to integrate the differential equations. The cost function, known as FF4 in the reference work as introduced, is implemented in Python (version 2.7.12) and linked to the NEST simulator. The simulation environment uses a common and fixed random-generation seed to define a stable framework and homogenize the evaluations. In this context, the GA method by Marín
et al. (
2020) was also implemented in Python 2.7.12 using the standard components provided by the DEAP library (Kim and Yoo,
2019).
Marín et al. configured their GA method to work with a population of 1000 individuals for 50 generations and selection tournaments of size 3. They adjusted the crossover, mutation, and per-component mutation probabilities to 60%, 10%, and 15%, respectively. Approximately, this configuration results in 30 000 evaluations of the objective function (f.e.) and also defines a general reference in computational effort. This aspect is especially relevant since the development and execution platforms differ from the original work. Namely, the four new optimizers compared (MemeGA, DE, TLBO, and MSASS) run in MATLAB 2018b after having wrapped the objective function of Marín et al. to be callable from it. The experimentation machine features an Intel Core i7-4790 processor with 4 cores and 32 GB of RAM.
The comparison takes into account four different computational costs in terms of the reference results: 50%, 75%, 100%, and 200% of the function evaluations consumed by GA, i.e. 15 000, 22 500, 30 000, and 60 000, respectively. Nevertheless, before launching the final experiments, the population-based optimizers have been tested under different preliminary configurations to find their most robust set of parameters. After having adjusted them, the focus was moved to those parameters directly affecting the overall computational cost, i.e. function evaluations. In practical terms, those are limited to the population sizes and the number of cycles for DE and TLBO. Similarly, they are the only parameters of the reference method, GA, that have been ultimately re-defined to encompass the new execution cases. However, for its memetic version, MemeGA, the number of objective function evaluations for each independent local search (local f.e.) has also been varied with the computational cost. In contrast to them, MSASS is directly configured in terms of function evaluations. Its local search component has kept the constant configuration explained below.
Table 2
Varied parameters for each population-based method and computational cost.
|
|
Computational cost (f.e.) |
| Method |
Parameters |
15k |
22.5k |
30k |
60k |
| GA |
Population |
1500 |
1500 |
1000 |
1000 |
|
Cycles |
16 |
25 |
50 |
100 |
| MemeGA |
Population |
500 |
500 |
300 |
300 |
|
Local f.e. |
10 |
12 |
20 |
35 |
|
Cycles |
20 |
26 |
40 |
50 |
| DE |
Population |
250 |
250 |
200 |
150 |
|
Cycles |
60 |
90 |
150 |
400 |
| TLBO |
Population |
250 |
250 |
200 |
150 |
|
Cycles |
30 |
45 |
75 |
200 |
Table
2 shows the previous information for each population-based optimizer and computational cost. Concerning the population sizes, the general paradigm followed opted for spawning larger populations with the lower computational costs to increase and speed up the exploration. More precisely, when the computational cost is below 100% of the reference method, the population is larger to accelerate the identification of promising regions in the search space and to compensate for the impossibility of allowing the optimizer to run more cycles. Numerically, the population sizes of GA remain in the range of the reference paper for the most demanding cases and get an increment of 50% for the others. Those of MemeGA come from GA but are approximately divided by 3 to allow for the function evaluations of its independent instances of SASS, which grow from 10 to 35 for the 60k configuration. The population sizes of DE are in the range of multiplying by ten the number of variables, as suggested by the authors, and doubled to maximize diversity. TLBO successfully shares the same strategy.
Regarding the number of cycles, it is adapted from the corresponding population size to achieve the desired computational effort. GA has statistical components, but its approximate number of function calls is 60% of the population size each cycle. MemeGA keeps the crossover and mutation probabilities of GA and redefines the latter as the percentage of individuals to be locally improved. Hence, it approximately executes 60% of the population size plus 10% of the referred value multiplied by the number of local evaluations each cycle. DE consumes as many f.e. as individuals per cycle, and TLBO takes twice. The cost of evaluating the initial populations is neglected.
Concerning the rest of the parameters that have been fixed, the internal SASS component of MemeGA considers
$\textit{Scnt}=5$,
$\textit{Fcnt}=3$,
$ex=2.0$, and
$c=0.5$ with
$\sigma \in \{1\mathrm{e}-5,0.25\}$. This is mainly the configuration recommended by Solis and Wets (
1981) with the only exception of the upper bound of sigma being 0.25 instead of 1. It is because preliminary experimentation revealed that when allowing few function evaluations, most individuals were directly moved to the bounds by big perturbations without the possibility to improve. Thus, the smaller upper bound of
σ allows for a better local exploration within MemeGA. However, for MSASS, all the parameters of SASS coincide with the recommended values because its execution approach does not have to share resources. That said, every instance of SASS will be stopped after 50 non-improving or failed iterations to save function evaluations for later independent starts. Regarding DE, after preliminary experimentation, it has been configured to use ‘rand’ selection, a single difference vector, and per-generation dither. Notice that the latter aspect avoids adjusting the
F parameter of the optimizer and takes into account its potential convergence advantages. Finally, the crossover rate has been fixed to 0.8, which is in the general-purpose range and between the values used in the extension proposed in Cabrera
et al. (
2011).
4.3 Discussion
From the results in Table
3, it is possible to make several overall appreciations. Firstly, none of the optimizers stably converges to a single global solution even after doubling the computational effort allowed in the reference paper. This aspect indicates that the search space is hard to explore and features multiple sub-optimal points. For this reason, ensuring stable and global convergence might not be feasible in a reasonable amount of time. Fortunately, it is not a requirement in this context as long as the found configurations make the simulated neurons behave as expected. Secondly, all the optimizers tend to improve the average quality of their results when the computational cost increases, but TLBO always achieves the best average quality. It is also the method that has found the best individual solution known so far, with a value of 87.45. Thirdly and last, the reduction in standard deviations is not as regular as that of the averages, but for most methods, the standard deviation of the results after 60 000 f.e. is approximately half of that observed for 15 000 f.e. The only exception is the reference optimizer, GA, whose STD was better for the lightest configuration than for the heaviest. Hence, in general, the probability of obtaining particularly divergent results in quality is reduced when the optimizers are provided with enough computational budget.
As previously commented, all the methods provide better average results after increasing the computational budget, but they effectively evolve at different rates. Certainly, the average quality achieved by a certain optimizer with a particular computational cost turns out to be nearly equivalent to that of another one. Nevertheless, some of the algorithms require more computational effort to be at the same level as others. For instance, the average of GA for 60 000 f.e. is very similar to those of MSASS and TLBO for 22 500 f.e. In fact, between 15 000 and 30 000 f.e., MSASS and TLBO are a step beyond the rest. Moreover, not all the methods ultimately achieve the same degree of quality. More specifically, for 60 000 f.e., MemeGA, DE and MSASS perform similarly, but GA is worse than them, and TLBO remains numerically ahead. Thus, according to the computational cost, some of the methods stand out from the rest. TLBO and MSASS do it positively with the best averages, and GA does it negatively considering the two heaviest configurations. Regarding MemeGA and DE, the former starts to outperform the reference method with 30 000 f.e., and the latter does it after doubling this value.
After the preliminary analysis, it is necessary to test whether the methods exhibit statistically significant differences considering the impact of uncertainty. For this reason, the individual results have been studied according to the Kruskal-Wallis test (Kruskal and Wallis,
1952; Mathworks,
2021), which is a non-parametric method for testing whether the samples come from the same distribution. By proceeding this way, it is possible to assess if the results registered for each optimizer and cost seem to significantly differ from each other with reduced datasets and without making distribution assumptions. The overall significance of the tests is 0.05, i.e. the corresponding confidence level is 95%.
Observing in depth the results in Table
3 for the lightest computational cost, it seems that there are two groups. Specifically, TLBO and MSASS exhibit the best performance with close values between them, while GA, MemeGA, and DE show worse averages (and also numerically similar between them). According to the Kruskal-Wallis test, there is no significant statistical difference within each group either. Thus, for 15 000 f.e., TLBO and MSASS return indistinguishable results within a significance of 0.05. In other words, for half of the computational budget of the reference work, TLBO and MSASS are equivalent according to the results achieved. Moreover, both perform better than the rest with a statistically significant difference, including the reference method, GA. Analogously, the performances of GA, MemeGA, and DE are equivalent in this context, so there is no more expected difference than the effect of uncertainty between the three. Additionally, the fact that the memetic variant of GA, MemeGA, does not significantly outperform it can be attributed to the lack of computational budget to let local search be effective.
For 22 500 f.e., the previous situation persists: TLBO and MSASS define the group of the best-performing optimizers, without significant difference between using one or the other. GA, MemeGA, and DE remain the worst-performing methods without practical difference between them. However, for 30 000 f.e., the situation changes: MemeGA moves from the group of GA and DE to that of TLBO and MSASS, which feature the best average. At this point, the methods achieving the worst results are GA and DE, which keep being statistically indistinguishable. Those with better performance are now TLBO, MSASS, and MemeGA. Among them, TLBO and MSASS keep being statistically indistinguishable within a significance of 0.05, but the same occurs between MemeGA and MSASS. Finally, the previous trend is confirmed with 60 000 f.e: another method, DE, separates from the reference one, GA, and outperforms it. Therefore, GA persists as the only member of worse-performing methods in the end, while MemeGA, DE, TLBO, and MSASS become statistically indistinguishable between them and achieve better results than the reference method.
Based on the previous analysis, TLBO and MSASS are the best choices for 50 and 75% of the function evaluations allowed in the reference paper. For the same computational budget considered by Marín
et al. (
2020), the performance of MemeGA reaches the level of those two referred methods, which remain ahead. Finally, with the highest computational cost, MemeGA, DE, TLBO, and MSASS significantly outperform the reference method, and there is no meaningful difference between the four. Thus, the reference method, GA, is always outperformed by at least TLBO and MSASS. MemeGA and DE also tend to separate from it to join the other two at 30 000 and 60 000 f.e., respectively.
Additionally, for the sake of completeness, the significance of the difference between the results of each method has also been systematically assessed with the Kruskal-Wallis test under the previous significance level. According to the study, there is no statistically significant difference between the results of GA after 22 500 and 30 000 f.e. Hence, it seems possible to reduce the computational effort in the reference work by 25% without expecting worse results for any cause apart from stochasticity. TLBO, MemeGA and DE experience the same phenomenon, but they do between 15 000 and 22 500 f.e, while the performance of the last two is still similar to that of GA. Finally, MSASS stagnates between 30 000 and 60 000 f.e., which is well aligned with its conceptual simplicity and lack of sophisticated components to globally converge.
4.4 Insight into the Best Solution
To conclude this section, the best result found will be analysed with further details in Fig.
2. The best-fitted neuron model according to all the features (with the lowest score) is an individual obtained by TLBO featuring a quality of 87.45 (blue lines). The neuron model from the reference work (GA optimizer) (Marín
et al.,
2020) with a score of 104.24 is also compared (orange lines). Finally, the
in vitro data used to define the fitness function are also represented (black dots) (D’Angelo
et al.,
2001; Masoli
et al.,
2017).
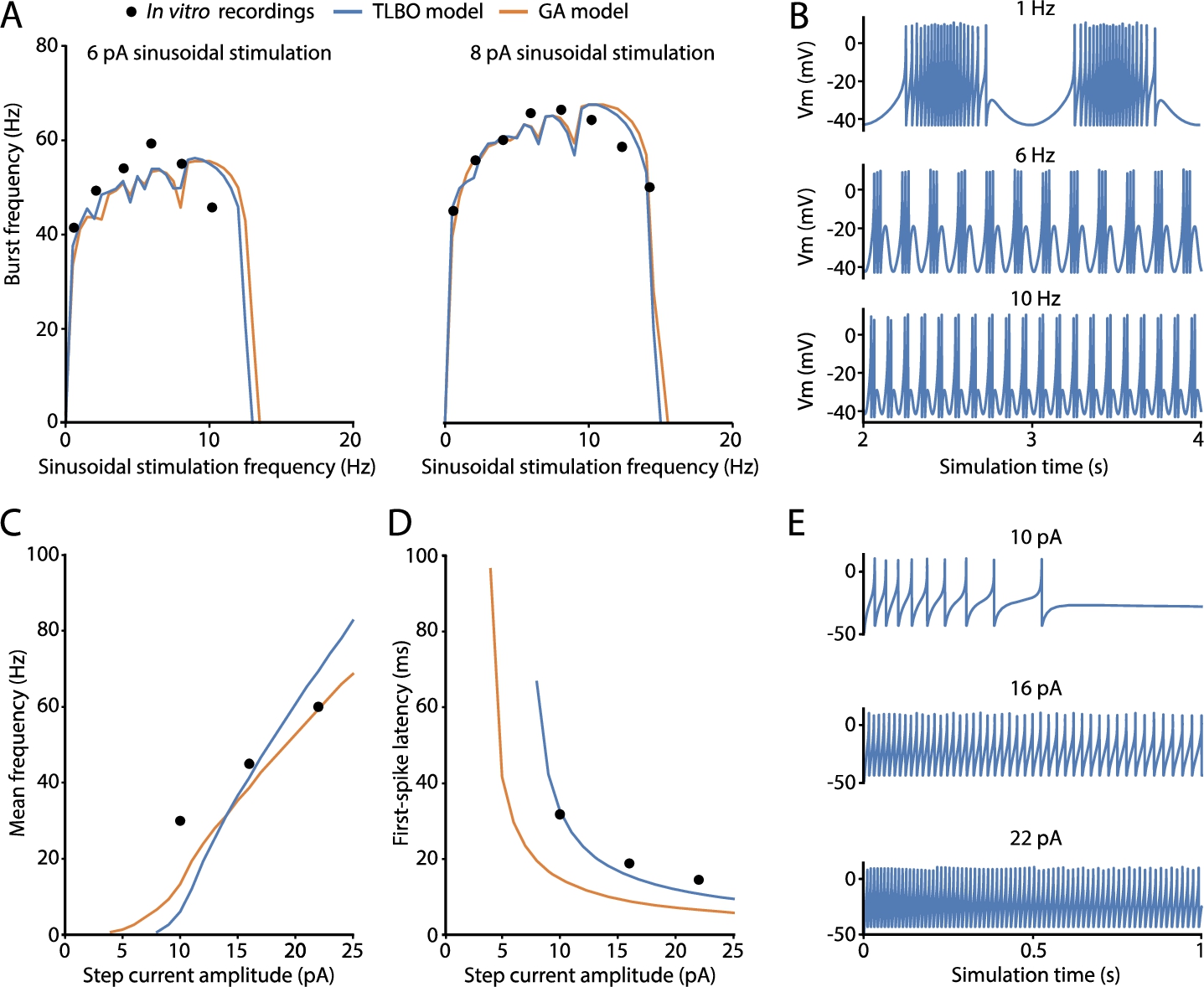
Fig. 2
Spiking properties predicted by the best-fitted cerebellar granule cell (GrC) model obtained by TLBO optimizer. Simulated features and electrophysiological traces of the best solution from TLBO (blue) and compared to the neuron model from the reference work (orange) and the target in vitro recordings used in the fitness function (black dots). A) Spiking resonance curves of the models computed as average burst frequencies in response to sinusoidal stimulation of 6 pA (left) and 8 pA (right) with increasing frequencies (in steps of 0.5 Hz). B) Membrane potential evolution of the TLBO model generates spike bursts in response to sinusoidal current injections with offset of 10 pA and amplitude of 6 pA. This is shown after 2 s of stimulation (stabilization). C) Repetitive spike discharge (intensity-frequency curves) of the models computed as the mean frequency in response to step-currents of 1 s. D) Latency to the first spike in response to step-currents of 1 s. E) Membrane potential traces of the TLBO model in response to step-current injections of 1 s with various amplitudes.
The selected neuron model has successfully captured the well-demonstrated features of the intrinsic excitability of cerebellar GrCs, i.e. repetitive spike discharge in response to injected currents (implemented as the mean frequency), latency to the first spike upon current injection (implemented as the time to the first spike), and spiking resonance in the theta-range (implemented as the average burst frequencies in response to sinusoidal stimulations). The neuron model resonates in the theta frequency band as expected, i.e. 8–12 Hz (Fig.
2(A)). The model practically reproduces identical resonance curves as the model of reference (GA model) (Marín
et al.,
2020). These resonance curves are the graphical representation of doublets, triplets, or longer bursts of spikes generated when stimulated by just-threshold sinusoidal stimulation at different frequencies (Fig.
2(B)). The main behaviour of biological GrCs is the increase of spike frequency when the latency to the first spike decreases as current injections increase. A sample of the neuron behaviour from which these features are calculated is shown in Fig.
2(E). The repetitive spike discharge of the TLBO model is similar to that of the model of reference and in accordance with the experimental measurements in real cells (Fig.
2(C)). The real improvement obtained by the neuron model of the proposed optimizer lies in the first-spike latency feature. The model from the reference work exhibited longer latencies than those experimentally reported, mainly with low stimulation currents (Fig.
2(D)). However, the TLBO model achieves an adjustment in its score of 7.95 ms compared to the 34.95 ms-error obtained by the GA model, both with respect to the
in vitro data. Thus, the TLBO optimizer proves not only to be effective in fitting the model parameters to diverse spiking features, but also to improve both the quantitative and qualitative predictions of these supra-threshold characteristics against the methodology of reference (GA).

